Login

RLIMS-P is a rule-based text-mining program specifically designed to extract protein phosphorylation information on protein kinase, substrate and phosphorylation sites from biomedical literature (Torii et al., 2015). RLIMS-P currently works on PubMed abstracts and open access full text articles.
We are currently working on enhancing the backend database for RLIMS-P. As a result RLIMS-P database is not being updated on a regular basis. Please bear with us.
RLIMS-P Search Form
Enter Keywords (accepts Boolean operators (AND, OR, NOT))
Or Enter PubMed IDs (PMIDs) delimited by "," or space, e.g., 15234272, 16436437.
You can process up to 200 PMIDs per run. Sample output
You can process up to 200 PMIDs per run. Sample output
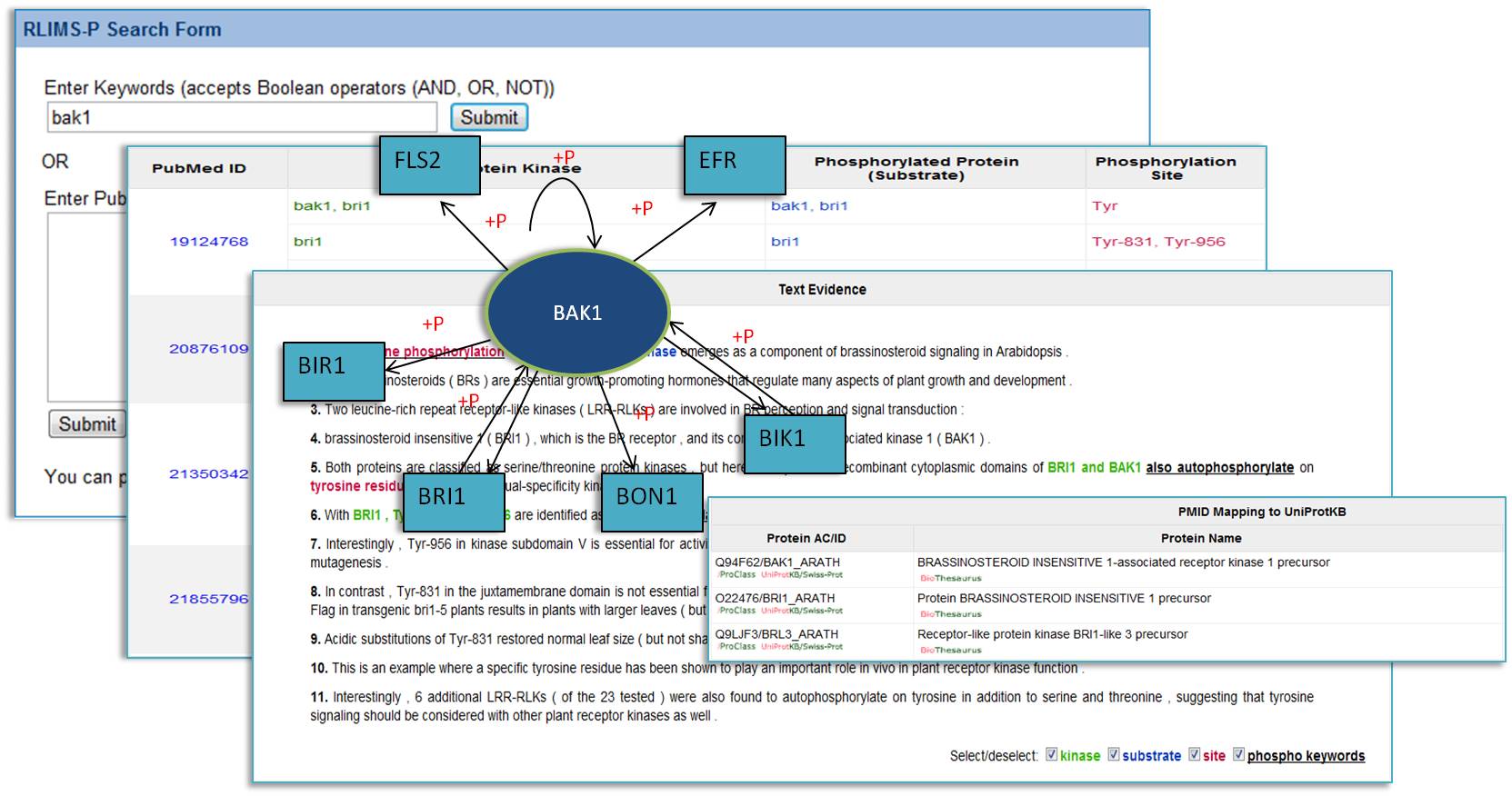
This RLIMSP website enables user search phosphorylation information by keywords or a list of PMIDs in the database. The results (kinase, substrate, site) are displayed in sortable tables with text evidence and downloadable for further research.